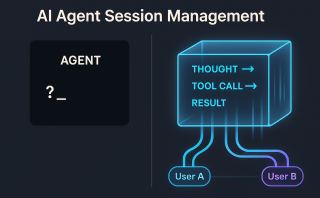
Part 9 of “My Journey in Building Agents from Scratch” — AI agent vector search with a YAML-driven ingestion pipeline
At the end of Part 8, I left a failure unresolved. A user asked for “that flexible screen phone” and the like "%flexible%" filter returned zero rows — because the Samsung Galaxy Z Fold 5 is described as a “foldable smartphone”, not a “flexible screen phone”. The words don’t match. The meaning does.
I knew this was coming. The moment I built the keyword filter tool, I could see it was a ceiling, not a solution. It worked perfectly for users who knew exact product names or categories. But real users don’t always know what something is called. They describe what it does, how it feels, what they’re trying to achieve. That gap — between a user’s words and a product’s data — is exactly what vector search exists to close.
This post is about how I built the ingestion pipeline that makes AI agent vector search possible: what decisions I made, what didn’t work, and why the final design looks the way it does.
Two Kinds of Data to Embed
My first instinct was to embed product records. Simple enough — one product, one vector. But as I thought about what the agent should ultimately know, I realised products were only half of it. Users also ask questions the product catalog can’t answer: “what’s the return policy for damaged items?”, “how long does express shipping take?”, “can I exchange a gift?” Those answers live in documents — policy files, FAQs, guides.
So I had two fundamentally different data shapes to deal with:
Structured data — product records from a database table. Each row is a self-contained piece of knowledge. One row, one vector. The challenge is deciding which columns to combine and in what narrative form.
Unstructured data — documents: policy files, product guides, FAQs. Each document is too long to embed as a single vector — the embedding model has a token limit, and a 4-page returns policy compressed into 384 numbers loses almost all specificity. The solution is chunking: split the document into overlapping windows, embed each chunk separately. One document, many vectors.
The ingestion logic for these two cases is different enough to warrant separate classes. But I wanted the same configuration pattern for both — declare the intent in YAML, let the class handle the mechanics.
The YAML Config: Declaring Intent, Not Code
I had already been using YAML to configure agents — the agent_type field, tool lists, session settings. It felt natural to extend that pattern here. Instead of hardcoding column names, table connections, and embedding logic into the ingestion script, I put them in a YAML file.
The core idea: the YAML declares what to embed. The Python class handles how.
This means onboarding a new data source never requires touching the Python code — just a new YAML file. I have seen what happens in teams when ingestion logic is spread across ten slightly different scripts. YAML prevents that drift.
Structured data config (products_ingestion.yaml):
db:
host: ${PGVECTOR_HOST}
port: ${PGVECTOR_PORT}
name: ${PGVECTOR_DB}
user: ${PGVECTOR_USER}
password: ${PGVECTOR_PASSWORD}
collection:
name: products
embed_columns:
- Name
- Category
- Description
embedding_logic: "The product {Name} belongs to {Category} -- {Description}"
Document config (policy_docs_ingestion.yaml):
db:
host: ${PGVECTOR_HOST}
port: ${PGVECTOR_PORT}
name: ${PGVECTOR_DB}
user: ${PGVECTOR_USER}
password: ${PGVECTOR_PASSWORD}
collection:
name: policy_docs
chunk_size: 500
chunk_overlap: 50
The field I spent the most time on was embedding_logic. My first version just concatenated the columns with pipes:
"Smartphones|Samsung Galaxy Z Fold 5|Foldable smartphone with Dynamic AMOLED display"
The search results were noticeably worse. Embedding models are trained on natural language — sentences, paragraphs, conversations. A pipe-separated string of field values is not natural language. Switching to a sentence template made a real difference:
"The product {Name} belongs to {Category} -- {Description}"
That reads like something a human might write. The model produces a better, more semantically rich vector from it.
All connection details come from environment variables. No credentials in YAML files, no credentials in code. The YAML marks the placeholders with ${ENV_VAR} and the processor resolves them at runtime.
The pgvector Schema
I already had a PostgreSQL database running — the same one holding order_view from Post 8. Adding pgvector was one line:
CREATE EXTENSION IF NOT EXISTS vector;
That’s it. No new service, no new infrastructure. The embeddings live in the same database as everything else, accessible via the same connection.
The schema is one table per collection, named from the YAML collection.name:
CREATE TABLE IF NOT EXISTS products (
id SERIAL PRIMARY KEY,
source_id TEXT NOT NULL, -- product_id or file path + chunk index
content_text TEXT NOT NULL, -- the text that was embedded (for debugging)
embedding vector(384) NOT NULL, -- all-MiniLM-L6-v2: 384 dimensions
metadata JSONB, -- any extra fields to return at search time
UNIQUE(source_id)
);
CREATE INDEX IF NOT EXISTS ON products
USING ivfflat (embedding vector_cosine_ops)
WITH (lists = 10);
The content_text column turned out to be more important than I expected. On my first real test, the agent was returning products that seemed wrong. I ran a raw SQL query and looked at the content_text values — and immediately saw the problem: I had been applying the embedding_logic template before stripping whitespace from the columns. Product descriptions with trailing newlines were producing slightly different vectors than clean ones. I would never have caught that without being able to inspect exactly what text went into each embedding.
Store the text. You will thank yourself later.
StructuredDataProcessor
My first pass at this was a standalone script — ingest_products.py. Read the product table, loop through the rows, embed each one, insert. It worked fine.
Then I needed to embed a second table — regional warehouse stock data, for a different collection. I copied the script, renamed the variables, and immediately hated what I was doing. The two scripts were 90% identical. Every fix to the embedding logic would need to be applied in two places. This is how ingestion code rots.
So I extracted a class. The logic stays the same; the configuration comes from YAML.
# structured_processor.py
# pip install sentence-transformers psycopg2-binary pandas pyyaml
import os
import re
import psycopg2
import pandas as pd
from sentence_transformers import SentenceTransformer
model = SentenceTransformer("all-MiniLM-L6-v2")
def resolve_env_vars(config: dict) -> dict:
"""Recursively replace ${VAR} placeholders with environment variable values."""
if isinstance(config, dict):
return {k: resolve_env_vars(v) for k, v in config.items()}
if isinstance(config, str):
return re.sub(
r"\$\{(\w+)\}",
lambda m: os.environ.get(m.group(1), m.group(0)),
config,
)
return config
class StructuredDataProcessor:
def __init__(self, df: pd.DataFrame, config: dict):
self.df = df
self.config = resolve_env_vars(config)
self.table = self.config["collection"]["name"]
self.logic = self.config["embedding_logic"]
self.cols = self.config["embed_columns"]
self._conn = None
def _get_conn(self):
if not self._conn or self._conn.closed:
db = self.config["db"]
self._conn = psycopg2.connect(
host=db["host"], port=db["port"], dbname=db["name"],
user=db["user"], password=db["password"],
)
return self._conn
def _build_text(self, row: pd.Series) -> str:
return self.logic.format(**{col: row[col] for col in self.cols})
def ingest(self):
texts = [self._build_text(row) for _, row in self.df.iterrows()]
embeddings = model.encode(texts, batch_size=64).tolist()
conn = self._get_conn()
cur = conn.cursor()
for (_, row), text, embedding in zip(self.df.iterrows(), texts, embeddings):
source_id = str(row.get("id") or row.iloc[0])
metadata = row.drop(labels=self.cols, errors="ignore").to_dict()
cur.execute(
f"""
INSERT INTO {self.table} (source_id, content_text, embedding, metadata)
VALUES (%s, %s, %s, %s)
ON CONFLICT (source_id) DO UPDATE
SET content_text = EXCLUDED.content_text,
embedding = EXCLUDED.embedding,
metadata = EXCLUDED.metadata
""",
(source_id, text, embedding, psycopg2.extras.Json(metadata)),
)
conn.commit()
cur.close()
print(f"Ingested {len(texts)} rows into '{self.table}'.")
Usage is two lines:
df = pd.read_sql("SELECT * FROM products", conn)
config = yaml.safe_load(open("products_ingestion.yaml"))
StructuredDataProcessor(df, config).ingest()
For the Samsung Galaxy Z Fold 5, _build_text() produces:
The product Samsung Galaxy Z Fold 5 belongs to Smartphones -- Foldable smartphone with Dynamic AMOLED display, 7.6-inch inner screen, S Pen support
That is what gets embedded. When a user later asks “flexible screen phone”, the query vector lands close to this vector because the model understands the conceptual overlap between “flexible screen” and “foldable AMOLED display”.
DocumentProcessor
Product rows were clean. Documents were not.
My first attempt at embedding the returns policy document was naive: read the whole file, embed it as one vector, store it. I ran a search query — “can I return a scratched screen protector?” — and got almost nothing useful back. The similarity scores were all in the 0.4-0.5 range. A 4-page document compressed into 384 numbers is too diluted. The model captures a kind of “topic average” of the whole document, not the specific clauses and conditions that matter for a real query.
The standard fix is chunking: split the document into overlapping windows, embed each chunk independently. When a user asks about screen protector returns, their query vector lands close to the specific chunk that discusses accessories — not a blurry average of the entire policy.
The overlap matters. If a key sentence falls exactly at a chunk boundary and gets split in half, neither chunk captures it well. Overlapping windows ensure every sentence sits fully inside at least one chunk.
# document_processor.py
import os
import re
import psycopg2
import psycopg2.extras
from pathlib import Path
from sentence_transformers import SentenceTransformer
model = SentenceTransformer("all-MiniLM-L6-v2")
class DocumentProcessor:
def __init__(self, file_path: str, config: dict):
self.file_path = file_path
self.config = resolve_env_vars(config) # same helper as above
self.table = self.config["collection"]["name"]
self.chunk_size = self.config.get("chunk_size", 500)
self.chunk_overlap = self.config.get("chunk_overlap", 50)
def _read_file(self) -> str:
return Path(self.file_path).read_text(encoding="utf-8")
def _chunk(self, text: str) -> list[str]:
chunks = []
start = 0
while start < len(text):
end = start + self.chunk_size
chunks.append(text[start:end])
start += self.chunk_size - self.chunk_overlap
return chunks
def ingest(self):
text = self._read_file()
chunks = self._chunk(text)
embeddings = model.encode(chunks, batch_size=64).tolist()
db = self.config["db"]
conn = psycopg2.connect(
host=db["host"], port=db["port"], dbname=db["name"],
user=db["user"], password=db["password"],
)
cur = conn.cursor()
for i, (chunk, embedding) in enumerate(zip(chunks, embeddings)):
source_id = f"{self.file_path}::chunk_{i}"
cur.execute(
f"""
INSERT INTO {self.table} (source_id, content_text, embedding, metadata)
VALUES (%s, %s, %s, %s)
ON CONFLICT (source_id) DO UPDATE
SET content_text = EXCLUDED.content_text,
embedding = EXCLUDED.embedding,
metadata = EXCLUDED.metadata
""",
(
source_id,
chunk,
embedding,
psycopg2.extras.Json({"file": self.file_path, "chunk": i}),
),
)
conn.commit()
cur.close()
print(f"Ingested {len(chunks)} chunks from '{self.file_path}' into '{self.table}'.")
The source_id uses the file path plus chunk index: "docs/returns_policy.txt::chunk_3". That means re-ingesting the file — after the policy is updated — cleanly replaces every chunk. No orphan vectors from the old version lingering in the table.
Usage mirrors the structured processor:
config = yaml.safe_load(open("policy_docs_ingestion.yaml"))
DocumentProcessor("docs/returns_policy.txt", config).ingest()
Fitting Into Your Existing Data Pipeline
One thing I kept thinking about while building this: ingestion can’t be a thing you remember to do. Manual steps get forgotten. A two-week-old product catalog that’s invisible to semantic search is worse than no semantic search at all — the agent silently fails and the user doesn’t know why.
Because both processors are plain classes, they slot into any existing data pipeline. New product added to the catalog? The same code path that inserts the row into the products table can call the processor in the same operation:
def add_product(product: dict):
# 1. Insert into the relational table as usual
db.execute("INSERT INTO products ...", product)
# 2. Ingest into pgvector in the same operation
df = pd.DataFrame([product])
config = yaml.safe_load(open("products_ingestion.yaml"))
StructuredDataProcessor(df, config).ingest()
The vector store stays in sync with the relational store — no separate job to schedule, no reminder to set. The same pattern works for documents: whenever a policy file is updated and uploaded, DocumentProcessor(file_path, config).ingest() runs as part of the same handler.
The AI Agent Vector Search Tool
With the embeddings in place, I needed a search endpoint. The first version was straightforward: embed the query, find the nearest vectors, return the results.
That first version had no similarity threshold. Nearest-neighbour search always returns results — even if nothing in the database is remotely related to the query. I found this out when someone asked “what’s the warranty on a cracked screen?” and the agent confidently cited a product description that mentioned “scratch-resistant coating” — a 0.49 similarity score that the agent treated as authoritative. That is a worse outcome than an honest “I don’t know.”
The threshold fixes this. Below 0.6, the endpoint returns an empty list. The agent receives no results and responds honestly rather than inventing an answer from a weak match.
# vector_search_api.py
# uvicorn vector_search_api:app --reload --port 8002
import os
import psycopg2
from fastapi import FastAPI, HTTPException
from pydantic import BaseModel
from sentence_transformers import SentenceTransformer
app = FastAPI()
model = SentenceTransformer("all-MiniLM-L6-v2")
def get_conn():
return psycopg2.connect(
host=os.environ["PGVECTOR_HOST"],
port=os.environ["PGVECTOR_PORT"],
dbname=os.environ["PGVECTOR_DB"],
user=os.environ["PGVECTOR_USER"],
password=os.environ["PGVECTOR_PASSWORD"],
)
class SearchRequest(BaseModel):
query: str
collection: str
limit: int = 5
threshold: float = 0.6
@app.post("/search")
def search_by_meaning(request: SearchRequest) -> list[dict]:
"""
AI agent vector search: find records by semantic meaning rather than exact keywords.
Use this tool when the user describes something by what it does, how it feels,
or its purpose -- rather than its exact name.
Examples:
query="flexible screen phone", collection="products"
query="what is the return policy", collection="policy_docs"
Returns: list of matches with content_text, metadata, and similarity score.
Low-confidence matches (similarity < threshold) are excluded.
"""
safe_limit = min(int(request.limit), 20)
query_embedding = model.encode(request.query).tolist()
# Validate collection name against an explicit allowlist.
# The collection name comes from the LLM -- never interpolate LLM output
# directly into SQL, even as a table name.
ALLOWED_COLLECTIONS = {"products", "policy_docs"}
if request.collection not in ALLOWED_COLLECTIONS:
raise HTTPException(status_code=400, detail=f"Unknown collection: {request.collection!r}")
conn = get_conn()
cur = conn.cursor()
cur.execute(
f"""
SELECT
source_id,
content_text,
metadata,
1 - (embedding <=> %s::vector) AS similarity
FROM {request.collection}
WHERE 1 - (embedding <=> %s::vector) >= %s
ORDER BY embedding <=> %s::vector
LIMIT %s
""",
(query_embedding, query_embedding, request.threshold, query_embedding, safe_limit),
)
rows = cur.fetchall()
cur.close()
conn.close()
return [
{
"source_id": row[0],
"content_text": row[1],
"metadata": row[2],
"similarity": round(row[3], 4),
}
for row in rows
]
The collection name injection check deserves a note. The collection parameter names the table to search. It comes from the agent — which means it ultimately comes from LLM output. LLM output must never go directly into a SQL query, including as a table name. I added an explicit allowlist and a 400 error for anything not on it.
The Samsung Fold Query — Resolved
The same query that produced zero results with like now works:
Query: “Show me orders for that flexible screen phone”
The agent calls search_by_meaning with query: "flexible screen phone", collection: "products":
[
{"source_id": "31", "product_name": "Samsung Galaxy Z Fold 5", "similarity": 0.89},
{"source_id": "28", "product_name": "Motorola Razr+", "similarity": 0.71}
]
The agent surfaces the top result, confirms with the user, then calls query_orders from Post 8 with the exact product name to retrieve the order history. Two tools, one query, no dead ends.
How the Agent Chooses Between the Two Tools
Adding vector search didn’t replace the structured query tool — it complemented it. The agent now has two tools for product questions, and the routing between them is guided entirely by docstrings.
| Tool | Use when |
|---|---|
query_orders(select_columns, filters) | User specifies known values: name, category, region, date, price |
search_by_meaning(query, collection) | User describes something by purpose, feel, or function |
The search_by_meaning docstring says: “use this when the user describes something by what it does rather than its exact name” — followed by concrete examples. Those examples are not documentation for human readers. They are the signal the LLM uses to classify an incoming query as intent-based. The more specific and realistic the examples, the more reliably the model routes correctly.
Pitfalls
1. Embedding Staleness
The class-in-pipeline approach solves daily staleness, but a bulk retroactive update — changing the embedding_logic template, or adding a new column to embed_columns — requires a full re-ingest of every record. A change to the sentence template changes all the vectors. Change the template, re-run the processor on the full dataset.
2. Model Consistency
The query must use the same model as ingestion. Switching from all-MiniLM-L6-v2 (384 dimensions) to all-mpnet-base-v2 (768 dimensions) makes all stored vectors incompatible — and the failures are silent. Results will just be wrong, not errored. Treat the model name as part of the schema. Store it in a config table and check at startup.
3. Chunk Size Tuning
For documents, chunk size is a retrieval quality parameter, not just a technical limit. Chunks too small lose context; chunks too large dilute the signal. 400-600 characters with 50-100 overlap is a reasonable starting range for most prose. Tune based on actual retrieval quality for your specific content.
4. Collection Name Injection
The collection parameter in the search request becomes a table name in SQL. Always validate against an explicit allowlist before interpolating. LLM output cannot be trusted for SQL interpolation, even for structural elements like table names.
Key Takeaways
- AI agent vector search closes the gap keyword matching cannot: user intent vs. data text that share meaning but not characters.
- The
embedding_logictemplate is a design decision, not just formatting. Natural language sentences produce better vectors than concatenated column values. StructuredDataProcessorandDocumentProcessorare plain classes because classes fit into pipelines. Ingestion can happen alongside the regular DB insert — no separate scheduled job.- First attempt at documents (embed the whole file) produced poor retrieval. Chunking with overlap fixed it. Every sentence needs to sit fully inside at least one chunk.
- The threshold matters. Without it, the agent confidently answers from a 0.49 match. An empty list is a better outcome than a wrong answer dressed as confidence.
- Validate collection names against an allowlist. LLM output must never go into SQL unvalidated, including table names.
- The query model and ingestion model must match. A silent model switch produces wrong results, not errors.
Try It Yourself
- [ ] Enable pgvector:
CREATE EXTENSION vector;— then create aproductstable with avector(384)column - [ ] Write a
products_ingestion.yamlwith anembedding_logictemplate and runStructuredDataProcessoron 20-30 product rows - [ ] Query directly in SQL:
SELECT content_text, 1 - (embedding <=> '[query vector]'::vector) AS sim FROM products ORDER BY sim DESC LIMIT 5 - [ ] Ask “flexible screen phone” — verify the Z Fold 5 appears; ask “noise cancelling headphones” — verify it finds products described as “active noise reduction”
- [ ] Try a query the catalog can’t answer well, observe the low similarity score, and add a threshold that suppresses weak results
- [ ] Embed a document with
chunk_size: 2000(no chunking effect) and compare retrieval quality againstchunk_size: 500 - [ ] Register
search_by_meaningandquery_ordersas agent tools and ask: “Show me last month’s orders for that foldable Samsung phone” — watch the ReAct loop call vector search first, thenquery_orderswith the resolved name
Conclusion
The Samsung Galaxy Z Fold 5 was always in the catalog. The agent just couldn’t find it — because like answers the wrong question. Vector search answers the right one: not “does this text contain these characters?” but “does this product match this concept?”
What made this work wasn’t the vector math — pgvector handles that. It was the decisions around the math: what text to embed, how to structure it, when to suppress a low-confidence result, how to keep the vectors in sync with the source data. Those decisions are where most of the real work happened.
The agent now has structured query, semantic search, and policy document retrieval as three separate tools. The next post steps back and looks at the full picture: six agent classes, YAML-driven instantiation, and the complete toolkit assembled into one deployable system.